genTB is a genomics tool for analyzing and predicting drug resistances to tuberculosis (TB).
The application allows for sharing, mapping, manipulating and linking TB genetic data to drug resistance phenotype as well as other epidemiological, demographic and host variables. The tool performs 3 main tasks in its current version:
Predict
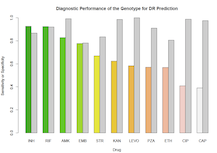
Clinicians and researchers can make predictions about an isolate’s resistance based on its genotype using a random forest classifier, trained using a 1400 isolate bank of clinical Mycobacterium tuberculosis. This classifier will be updated and retrained with the contribution of new high quality genotype and drug resistance data from the research community.
Data
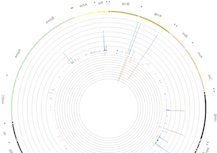
Clinicians and researchers can also store and share TB genetic and phenotypic data, allowing for the expansion of the current dataset. Every additional dataset that is uploaded is given a persistent and citable identifier using the broadly supported Digital Object Identifier (DOI) system, an ISO standard which provides an actionable, interpretable, and persistent link.
Map
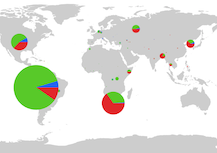
The tool also makes mapping TB genetic and phenotypic data as well as resistances easy. In particular, the tool, as it stands, allows for the mapping of resistance frequencies by different geographies through an interactive interface.
In Development
Stay tuned for additional analysis features as we have several in progress. If you would like to receive updates on these features please email us at tbpredict@gmail.com.
Training Data Sources
Farhat MR et. al. Genomic analysis identifies targets of convergent positive selection in drug-resistant Mycobacterium tuberculosis Nat Genet 2013
Farhat MR et. al. Genetic determinants of drug resistance in Mycobacterium tuberculosis and their diagnostic value AJRCCM 2016
Groeschel M et. al. GenTB: A user-friendly genome-based predictor for tuberculosis resistance powered by machine learning Genome Medicine 2021
Random Forest Classifier
Development and validation of this Prediction classifier is described in detail in:
Farhat MR, Razvan S et. al. Genetic determinants of drug resistance in Mycobacterium tuberculosis and their diagnostic value AJRCCM 2016
Groeschel M et. al. GenTB: A user-friendly genome-based predictor for tuberculosis resistance powered by machine learning Genome Medicine 2021
Terms of Use
This project is developed and maintained by researchers and clinicians as a benefit to the research and education community. Although this website is curated, it is also provided in an "as is" basis only, without warranty or representation as to its accuracy or reliability.
Cite the Project
Groeschel M et. al. GenTB: A user-friendly genome-based predictor for tuberculosis resistance powered by machine learning Genome Medicine 2021

